Welcome to the NEST Simulator documentation¶
NEST is used in computational neuroscience to model and study behavior of large networks of neurons.
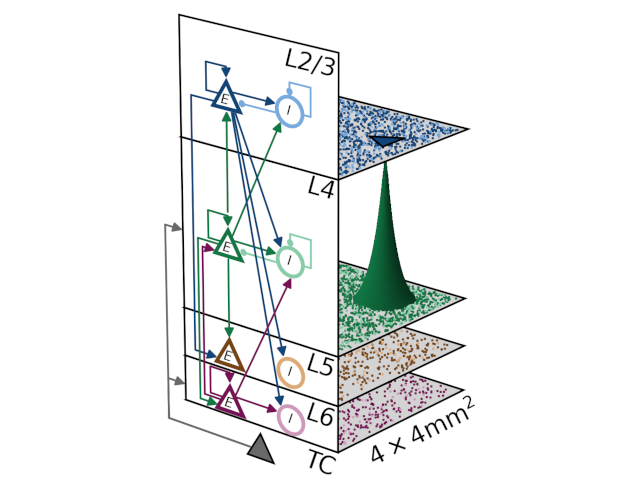
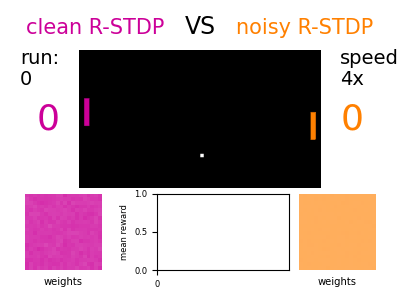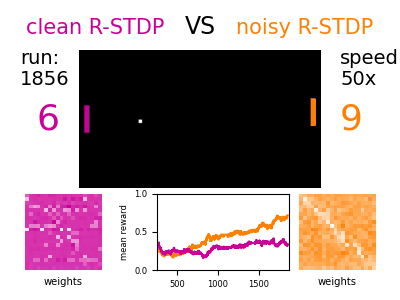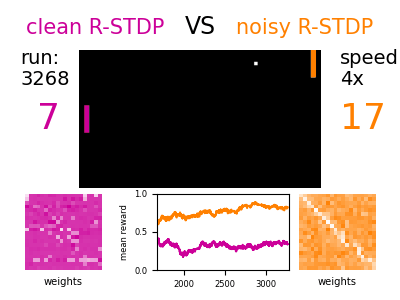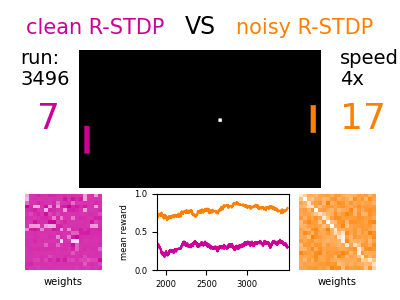
The models describe single neuron and synapse behavior and their connections. Different mechanisms of plasticity can be used to investigate artificial learning and help to shed light on the fundamental principles of how the brain works.
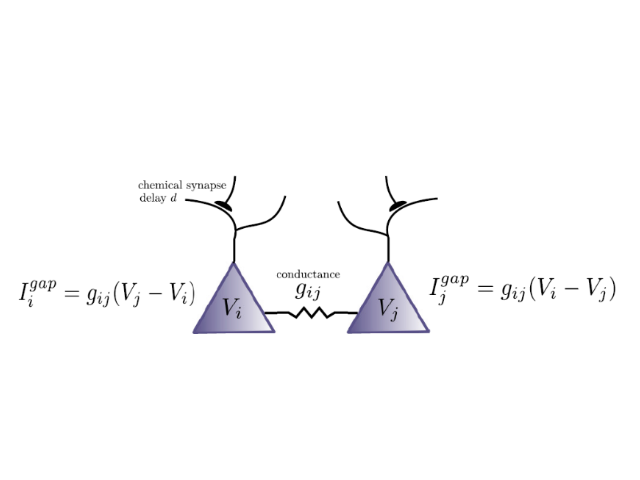
NEST offers convenient and efficient commands to define and connect large networks, ranging from algorithmically determined connections to data-driven connectivity. Create connections between neurons using numerous synapse models from STDP to gap junctions.